An interdisciplinary BBSRC Strategically Longer and Larger (sLoLa) project expanding the frontiers of protein-based biological circuitry




Creating and comprehending the circuitry of life: precise biomolecular design of multi-centre redox enzymes for a synthetic metabolism

The Project
A defining characteristic of life is the requirement of energy from an external source; we eat, plants absorb light. To maximize the energy gained from the food that we and all oxygen-breathing organisms consume, oxygen is converted to water as a final step and carbon dioxide is released. The oxygen in this equation arises from plants as they convert water, carbon dioxide and light, into oxygen and fuel. This cycle is not merely an auspicious result of billions of years of evolution. The molecular events that allow the processes of respiration and photosynthesis to happen are connected in deep ways, down to shared structures, molecules, and mechanisms.
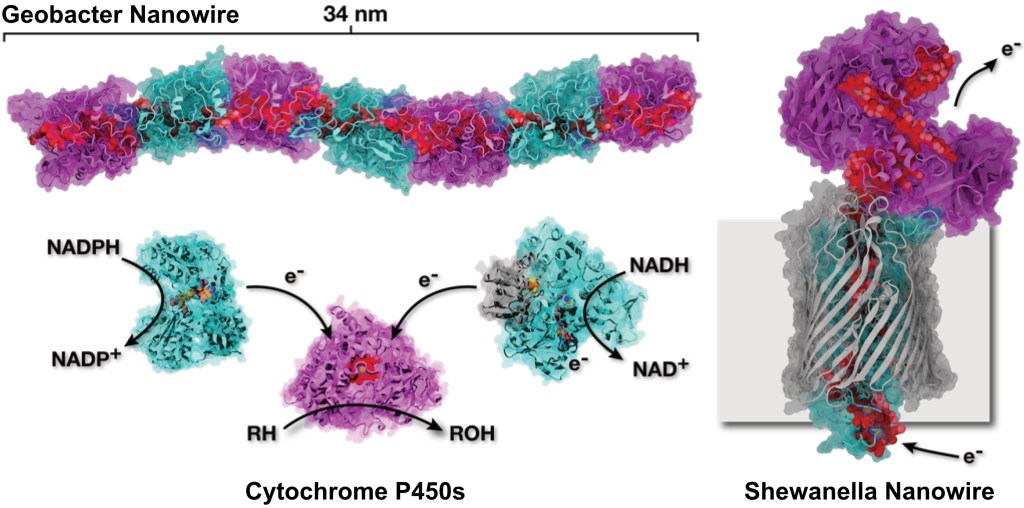
At their most basic, respiration and photosynthesis are Nature’s way to capture and convert energy from one form to another. To do this, Nature has evolved complex structures, termed oxidoreductases, that bind molecules that aid in this conversion. These molecules can both absorb light, imparting plants with their colours, and take and give electrons. The oxidoreductases have evolved to take energy from external sources and convert it into forms that can be used by living organisms to grow and survive. The evident complexity of this process belies a central feature of the oxidoreductases involved: evolution has yielded structures that are built from repeats of relatively simple modules. All of respiration and photosynthesis are built on these repeating modules. But despite nearly a century of investigation, where we have outlined how respiration and photosynthesis work in fine detail, we remain unable to construct our own models of these processes. This naturally leads to a question of whether we really understand how these processes occur.
Here we have assembled a team of researchers from multiple academic institutions and disciplines to address deficiencies in our knowledge, with the unified target of building completely new oxidoreductases from scratch. Through this work we will fill holes in our understanding of how Nature captures and converts energy.
Our work begins by combining powerful computational techniques that allow us to design and construct oxidoreductases with tailor made functions. Within a virtual reality framework that we are developing for this project, we will work together in a shared digital space to construct molecular binding sites, alter how molecules take and give electrons or catalyse reactions, and create oxidoreductase modules that, taking inspiration from Nature, we will join to produce more complex functions. With these designs, we will use an iterative ‘build-test-learn’ approach to construct new oxidoreductases that match the activities and actions of those Nature uses in respiration and photosynthesis. By pulling together our expertise in computational biophysical methods, oxidoreductase engineering, modular structure creation, molecular binding site assembly and their chemistry, and the analysis of very fast oxidoreductase functions, our team stands to make a substantial leap in our understanding of how to construct new oxidoreductases that has, so far, remained beyond our grasp.
The principles we establish through this work will help us to better understand the oxidoreductases of respiration and photosynthesis, finally clarifying architectural features that are essential for their assembly and function that have remained opaque for over a century. With our new sets of design principles, we will be able to create oxidoreductases that fulfil our needs in bioscience and biotechnology, from the creation of single structures that produce fuels from light, water and carbon dioxide akin to photosynthesis to biosensors that detect toxins in the environment or signs of disease.
Recent Posts
-
Two Reviews published!
The first is a perspective of our grand goals in building bioenergetic proteins and circuits from scratch, and an overview of our current de novo modules. Importantly, this review gives a summary of what we think are the main outstanding challenges. Read more here: Building tailor-made bioenergetic proteins and circuits from de novo redox proteins The second…
-
Single-heme de novo proteins enable the dissection of redox cooperativity in diheme cytochromes
We recently reported our work constructing single-heme proteins from our water-soluble and transmembrane diheme proteins 4D2 and CytbX. Whilst we initially set out to make these proteins for ‘completionist’ reasons, we realised we could use them to gather valuable insights to the factors that lead to redox potential splitting in our diheme proteins. Read the…
-
Fluctuation Relations to Calculate Protein Redox Potentials from Molecular Dynamics Simulations
We are excited to share a new tool, dubbed MD+CB, to calculate protein redox potentials based on fluctuation relations extracted from molecular dynamics simulations of heme proteins Spearheaded by Sofia Oliveira, Ross Anderson and Adrian Mullholland from the Circuits of Life project, this novel method was tested on point mutants (designed by past PhD student…
-
Congratulations to Ethan Bungay! Poster Prize winner
Congratulations to PhD student Ethan Bungay for winning the Best Poster Prize at this year’s Christmas Bioenergetics Meeting in York – well deserved!
-
4D2 Paper: Press-release from the University of Bristol
Pioneering study signals new era of environment-friendly programmable bioelectronics The University of Bristol have published a press release on our recent publication describing the design of 4D2, m4D2 and e4D2 “While our designs take inspiration from the protein-based electronic circuits necessary for all life on Earth, they are free from much of the complexity and…